Despite various rounds of testing before novel drugs come on the market, recalling occurs often. In the last nine years, 552 products were recalled in the US due to incorrect strength of the product, 2110 due to product quality and 701 due to stability failure in vivo [1]. Some products were even found to cause previously unknown toxicities to one or more organs, leading to death in a few cases [2]. Fortunately, advances in tissue engineering have led to the development of organoids, which are small three-dimensional cell cultures mimicking organs in vitro.
Organoids have been found to be able to resemble large in vivo organs better than 2D cell cultures, due to a more physiologically relevant representation of cell-cell interactions, genotype and phenotype [2]. These properties greatly influence the parameters that have to be evaluated in the drug development process, such as cytotoxicity, dosage dependency, pharmacodynamics and activity of the drug in various cell types [3]. Organoids are therefore widely used for studying bacterial and viral infections [4][5], disease modeling in cancer drug development [6][7], and drug toxicity assays [8], among others.
Organoids are often used to screen the toxicity of drugs in specific tissues.
This is especially important for toxicity to the liver. Drug-induced liver injury is a common cause of liver failure, but the toxicity of a drug to the liver is often unpredictable [9]. In liver organoids, drug toxicity can be determined by investigating the enzyme activity and the structure of liver cells.
Organoids are also used to screen the toxicity in drugs containing nanomaterials. These drugs offer great advantages, but often have unknown toxicities. Nanodrug toxicity studies have therefore been done in various organoids, such as the liver, kidney, brain and lungs [10]. The interaction of the nanodrug with the organs can then be closely monitored, as well as the accumulation of the drug in an organ.
To accurately characterize the organoid structure and the localization of drugs, a high-resolution imaging technique is crucial.
Imaging is therefore often performed using electron microscopy (EM), which makes it possible to view the ultrastructure of the organoid [4][11]. 2D EM however lacks volumetric information. To accurately view the whole organoid in high resolution, researchers use volume electron microscopy (volume-EM) and obtain 3D EM images.
In the volume-EM imaging workflow, the sample is usually embedded in resin. The resin-embedded sample is then cut into small slices, often called ribbons, and these are imaged using an electron microscope. After imaging, a 3D reconstruction can be made from the 2D images. These reconstructions show the ultrastructure of the whole organoid in 3D. This could be used to study:
- Detailed cell-cell interactions
- The ultrastructural morphology of the organoid
- Precise localization of the drugs.
Recently, Volume-EM imaging of an organoid was used to study bacteria causing urinary tract infections in the bladder [12].
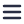
The researchers imaged a bladder organoid, in which they injected bacteria. They made a 3D map of the whole organoid, and enabled the visualization of the individual bacteria and the various cell types (Figure 1). In this way, they could identify five bacterial subpopulations based on their location and investigate their different properties quantitatively. They found that some invading subpopulations are protected from clearance by antibiotics.
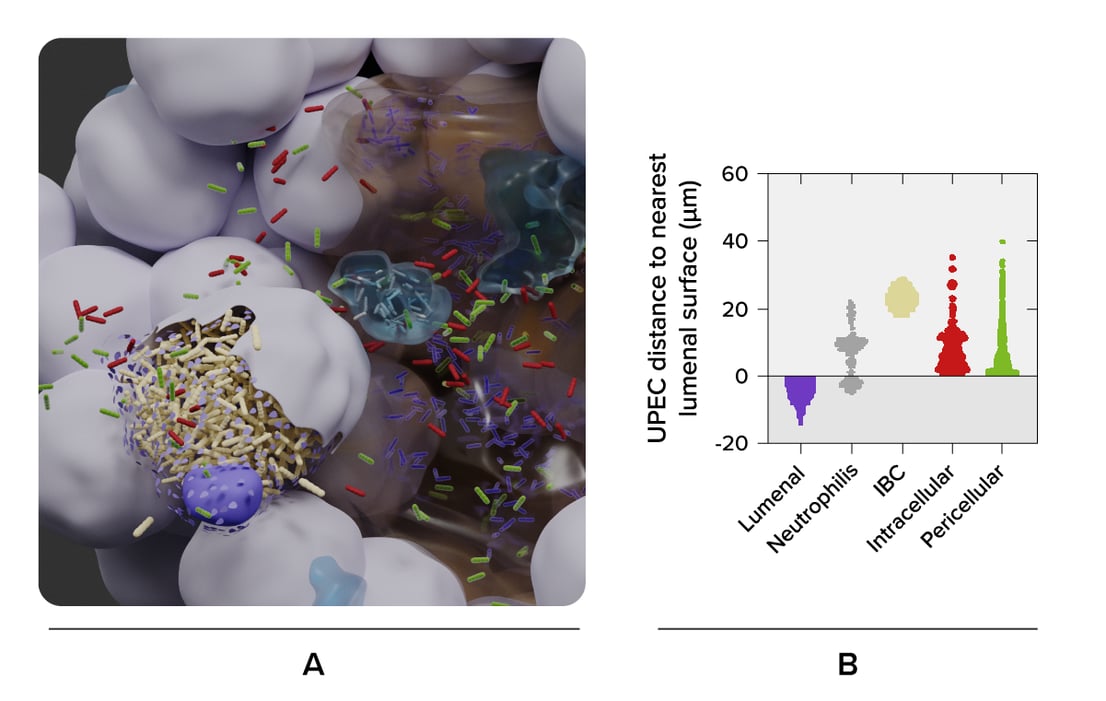
Figure 1: 3D reconstruction of a bladder organoid invaded by bacteria using Volume-EM. A) part of the organoid showing the lumen (brown), neutrophils (cyan) and epithelial cells (gray). Five bacterial subspecies can be seen colored violet (lumenal), gray (neutrophil), yellow (IBC), red (intracellular), and green (pericellular). B) Distances of the five bacterial subspecies to nearest luminal surface. Using the 3D model, quantitative analyses can be performed.
In another study by Ronchi et al. [13], researchers optimized the volume-EM workflow for imaging organoids among others. They imaged a mouse mammary gland organoid successfully and could identify the single cells in a 3D map.
Despite these great achievements, volume-EM has only been scarcely used for drug development and drug toxicity studies on organoids.
This is mainly because volume electron microscopy is not yet broadly accessible for researchers. The challenges of conventional scanning electron microscopes (SEMs) and the volume-EM workflows are that image acquisition is very slow, and working with these large datasets is challenging.
To make volume-EM more accessible and faster, Delmic developed the innovative FAST-EM. This is an advanced scanning transmission electron microscope that images up to 100 times faster than the conventional SEM. Its substrate contains a metal coating that prevents the sample from charging, thus omitting the coating step to make the samples conductive.
Using the FAST-EM, researchers can image large volumes, such as organoids, in a fast and automated way. The acquired images are automatically digitized and can be easily reviewed because of the innovative data management. These 3D high-resolution images could be used to review the toxicity and other effects of drugs on organoids in a fast manner.
To conclude, organoids are crucial for screening the effect of drugs on organs. Studies performing volume-EM imaging on organoids show a great level of detail and numerous novel biological insights that can be gained. Using FAST-EM, volume-EM can become an advanced high-throughput imaging technique. This could speed up the drug development process, therefore increasing the amount of drugs on the market that are effective and safe.
References
[1] Ashlee et al., Journal of the American Pharmacists Association 62, 4, 1344-1350, (2022)
[2] Devarasetty et al., BioDrugs 32, 53-68, (2018)
[3] Andrade et al., Brazilian Journal of Medical and Biological Research 49, 11, (2016)
[4] Barreras et al., J. Neurovirol. 28, 17–26, (2022)
[5] Puschhof et al., Nat Protoc 16, 4633–4649, (2021)
[6] Seidlitz et al., Gut 68, 207–217, (2019)
[7] Pauli et al., Cancer Discov 1, (2017)
[8] Xu et al., Exp Hematol Oncol 7, 30, (2018)
[9] Brooks et al., Pharmacological Research 169, (2021)
[10] Nabi et al., Journal of Applied Toxicology 42(1), 52– 72, (2022)
[11] Lee et al., Int. J. Mol. Sci. 21, 2982, (2020)
[12] Sharma et al., Cell Reports 36 (3), (2021)
[13] Ronchi et al., J Cell Biol 220 (9), (2021)
.png)