The authors present an imaging workflow that allows obtaining correlative super-resolution light and scanning electron microscopy images in an easy and fast way.
Correlative light and electron microscopy (CLEM) has been proven to be a powerful method for obtaining both structural information and chemical specificity with electron microscopy and light microscopy. This method is particularly useful for studying biological processes and structures. However, most of the methods for obtaining such information are complex and difficult to implement since they include multiple sample transfers and specific algorithms for image alignment and correlation. The paper presents the advantages of using the SECOM system, an integrated correlative light and electron microscope, for performing simple and cost-effective superresolution CLEM. The authors applied a recently developed algorithm called Super Resolution Radial Fluctuations (SRRF) for superresolution imaging with the standard wide field optical system of the SECOM, and the image analysis was performed using only an ImageJ plugin.
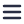
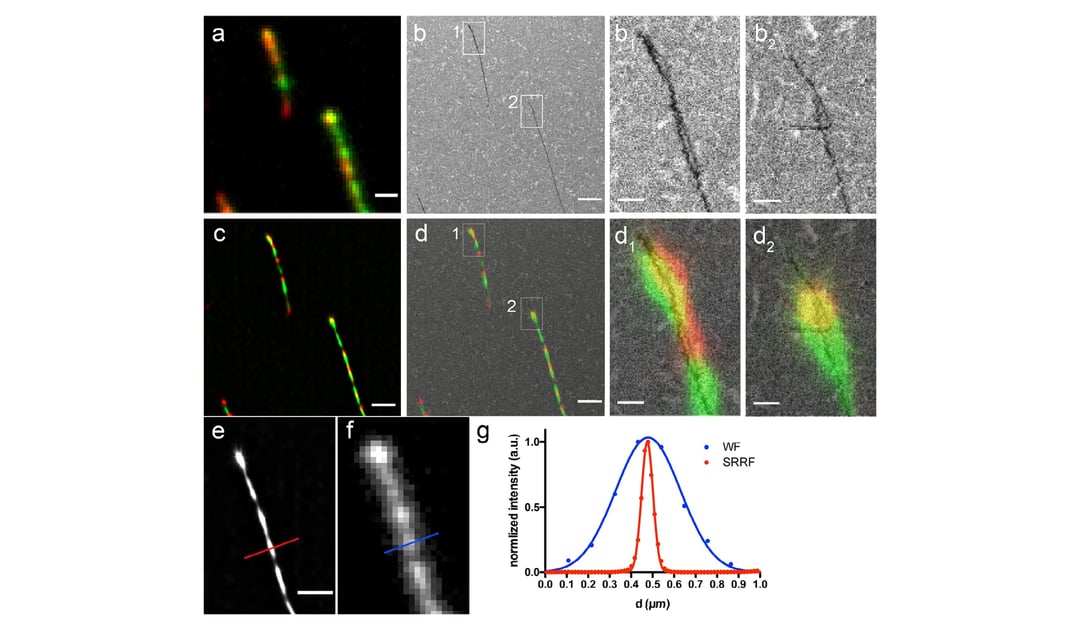
To demonstrate the value and potential of this method, the morphology of dual-color labeled amyloid fibrils formed by the human α-Synuclein (α-Syn) protein was studied. Their aggregation is connected to the onset of Parkinson’s disease, so studying these amyloid fibrils could be highly interesting. Moreover, non-pathological amyloid-like fibrils can be formed in vitro from small peptides and used for technological purposes. With the methodology proposed by the authors it was possible to easily discern details of the amyloid fibril aggregation and morphology. The authors demonstrate that this workflow is faster and easier than other methods, and could be extremely beneficial for research on amyloid protein aggregation kinetics.
The paper provides details into the all the methods that were used to achieve the results. To read the whole paper, please visit the page of the journal.
.png)