Over the last decade, there has been a boost in small-scale life science research. Advancements in electron microscopy (EM) technologies have enabled researchers to visualize the biological world of the small, leading to an enhanced understanding of cellular processes, human diseases, and viral infections. Using this knowledge, researchers have been able to develop novel treatments and vaccines at an accelerated speed.
However, the world of the small occurs in three dimensions, and for a long time, EM imaging of biological samples was only done in two dimensions. This limited our ability to fully understand cellular architecture, tissue organization, and dynamic interactions between cellular components. To address the need for high-resolution imaging of volumes, 3D or volume electron microscopy techniques were developed.
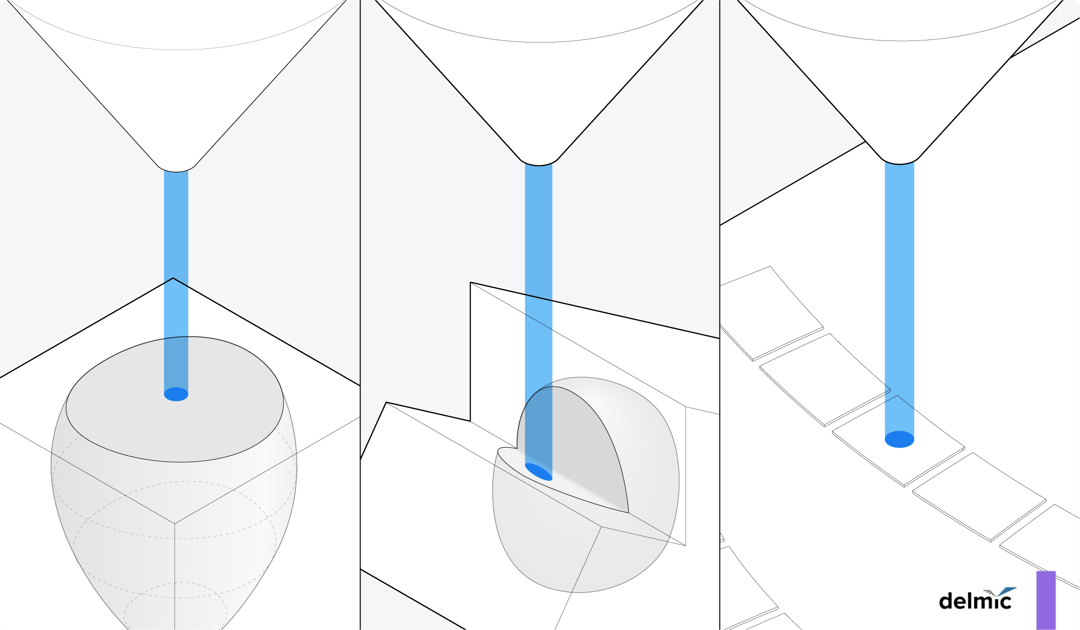
There are three main volume electron microscopy techniques that use a scanning electron microscope (SEM) for imaging: focused ion beam SEM (FIB-SEM), serial block face SEM (SBF-SEM), and serial section array tomography (AT).
All these techniques typically require tissue samples to be 1) fixed, 2) stained with heavy metals to increase contrast, 3) dehydrated, 4) resin infiltrated, and 5) resin embedded (Figure 1) [1]. The main differences between the 3D EM techniques lie in how the resin-embedded sample is subsequently cut and imaged using the SEM. Below, we will explain each technique in detail.
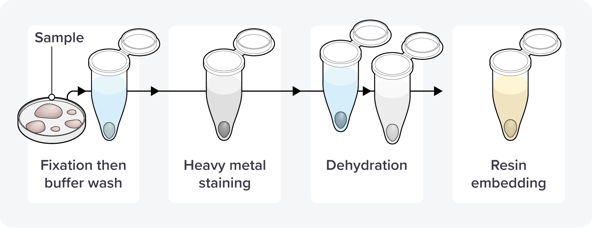 Figure 1: The first steps in the volume or 3D electron microscopy workflow, including fixation, staining, dehydration, and resin embedding.
Figure 1: The first steps in the volume or 3D electron microscopy workflow, including fixation, staining, dehydration, and resin embedding.
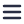
In focused ion beam SEM (FIB-SEM), layers of the resin-embedded sample are removed by a focused ion beam within an SEM. Typically, a minimum of 10 nm of the resin block surface is milled away, followed by SEM imaging of the newly exposed surface. These steps are iteratively executed until a z-stack of 2D images covering the entire sample is acquired.
The primary advantage of FIB-SEM, in comparison to other vEM techniques, is the superior z-resolution. However, the technique is inherently destructive to the sample, leaving no room for error during the milling and imaging processes. Additionally, the depth of tissue that can be effectively imaged is limited to a maximum of 1 mm, and the imaging process is relatively slow [1][2].
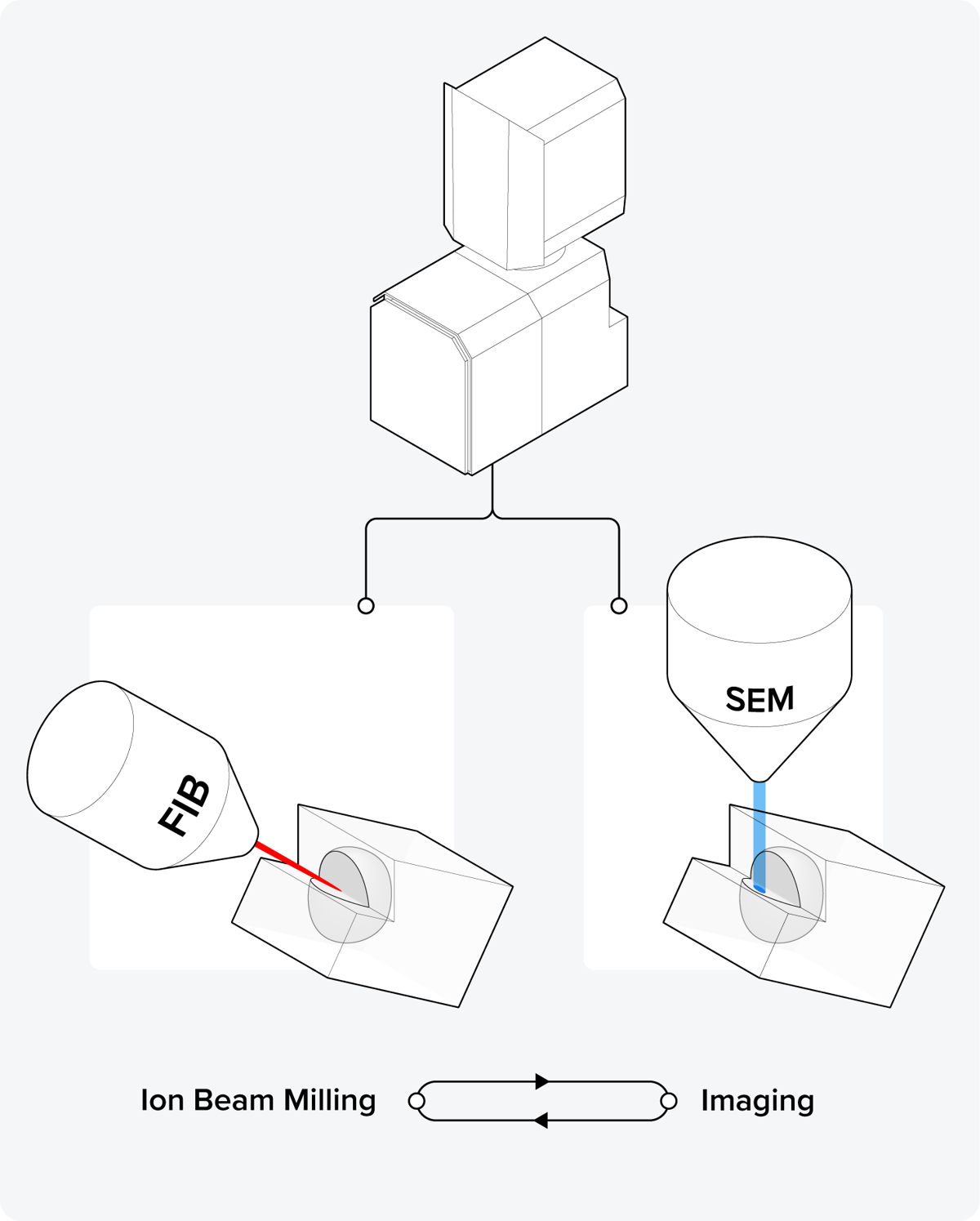 Figure 2: The principle of focused ion beam SEM (FIB-SEM). Inside the SEM, iterative steps are executed where a layer of the resin block surface is milled away by a focused ion beam, followed by SEM imaging of the surface.
Figure 2: The principle of focused ion beam SEM (FIB-SEM). Inside the SEM, iterative steps are executed where a layer of the resin block surface is milled away by a focused ion beam, followed by SEM imaging of the surface.
In serial blockface SEM (SBF-SEM), a microtome equipped with a diamond knife is located inside the vacuum chamber of the SEM. The process involves iteratively slicing thin sections from the resin-embedded sample and imaging the newly exposed surface of the block face. These repetitive cycles result in the acquisition of a z-stack composed of image slices, and the process continues until the entire sample has been cut and imaged. The cutting and imaging of sections are typically automated, allowing the microscope to operate unattended during the imaging process.
However, in SBF-SEM, artifacts can arise and the cellular structure of the sample can be altered during the cutting process. Consequently, drawing accurate conclusions based on the imaging results might be compromised. Additionally, there is a risk of electron accumulation on the resin block, potentially leading to sample damage, charging, and loss of contrast. Charging and damage can be mitigated with appropriate sample preparation for biological tissue and using a focal charge compensation device [3][4].
Lastly, similarly to FIB/SEM, the technique is inherently destructive. There is only a single opportunity to capture an image of a particular section. If the user wishes to inspect the same area again at a higher resolution or with a parallel technique, it is not possible anymore.
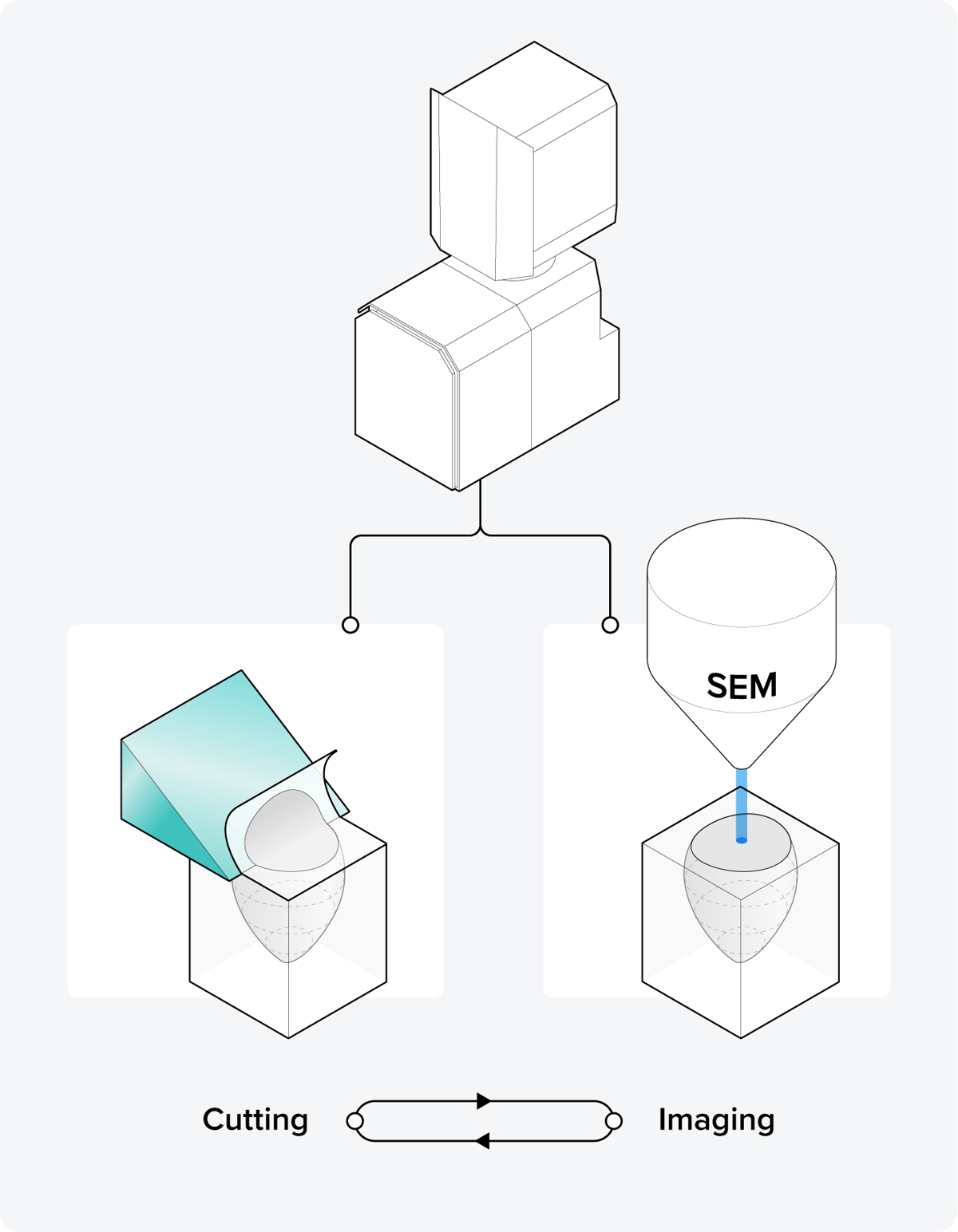 Figure 3: The principle of serial blockface SEM (SBF-SEM). Inside the SEM, iterative steps are executed where the top layer of the resin block surface is cut by a microtome equipped with a diamond knife, followed by SEM imaging of the surface.
Figure 3: The principle of serial blockface SEM (SBF-SEM). Inside the SEM, iterative steps are executed where the top layer of the resin block surface is cut by a microtome equipped with a diamond knife, followed by SEM imaging of the surface.
In serial section array tomography (AT), an ultramicrotome is used to cut the resin-embedded sample into small, thin sections. The sections can take the form of individual serial sections, or be arranged in arrays of sections known as ribbons. Subsequently, these sections are collected on a substrate for SEM imaging. There are various methods for section collection on the wafer, and researchers have invented some section collection tricks to facilitate this process.
Array tomography presents some challenges, including sections that could occasionally get lost, leading to an incomplete 3D image, and aligning the 2D images can be more complex compared to those obtained through the previously mentioned techniques. However, the significant advantage of array tomography over alternative techniques lies in its non-destructive nature, allowing for storage and reimaging of sections. Moreover, it can cut the widest sections, as wide as 4 mm [1].
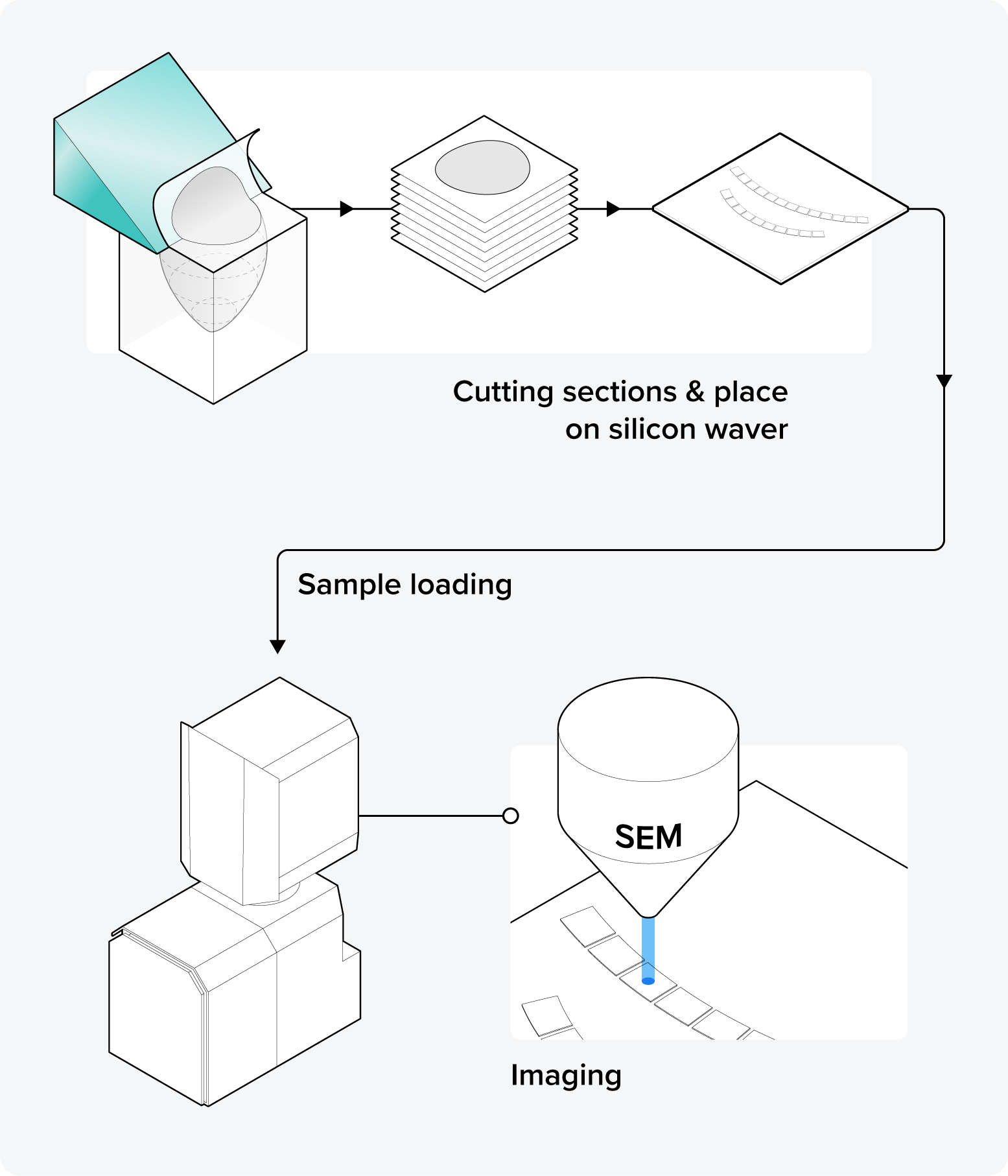 Figure 4: The principle of serial section array tomography (AT). The resin block is first cut into sections using an ultramicrotome. Then, the sections are loaded into the SEM and individually imaged.
Figure 4: The principle of serial section array tomography (AT). The resin block is first cut into sections using an ultramicrotome. Then, the sections are loaded into the SEM and individually imaged.
So which volume electron microscopy technique is the best?
There isn’t one superior volume-EM technique; each one has a trade-off between factors such as size, resolution, completeness, and usability. The choice depends entirely on the specific research question of the user. Furthermore, these techniques are continuously evolving, with ongoing innovations that aim to enhance their capabilities [5].
The primary focus of recent innovations is focused on improving the z-resolution of these techniques. However, there is an ongoing debate about the significance of a high z-resolution, especially in the field of connectomics [6]. Another area of improvement is the automation of the cutting and collection processes in array tomography, aiming to reduce the risk of losing sections and ending up with incomplete datasets.
A significant challenge in volume-EM is the image acquisition speed, as conventional SEMs have only one electron beam to scan a section or surface. Recent developments address this challenge, such as Delmic’s FAST-EM microscope, which contains 64 electron beams working in parallel and therefore vastly improves the acquisition speed.
In summary, the main 3D or volume electron microscopy techniques include FIB-SEM, SBF-SEM, and array tomography. The selection of these techniques depends on the specific research question. As ongoing innovations continue to enhance each technique, their trade-offs might evolve over time, potentially leading to the dominance of one technique in the future.
Curious about what the future of volume electron microscopy might look like? Watch our video series below to find out!
References
[1] Laws, R. et al., Journal of Investigative Dermatology 142, 2, 265-271 (2022)
[2] Smith, D. et al., Tissue and Cell 57, 111-122 (2019)
[3] Courson, J. A. et al., J. Vis. Exp. 169, e62045, (2021)
[4] Deerinck, T.J. et al., Journal of Microscopy, 270, 142-149 (2018)
[5] Kievits, A. J. et al., Journal of Microscopy 287, 114–137 (2022)
[6] Briggman, K.L. et al., Current Opinion in Neurobiology 22, 1, 154-161 (2012)
.png)