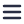
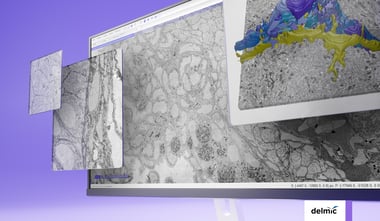
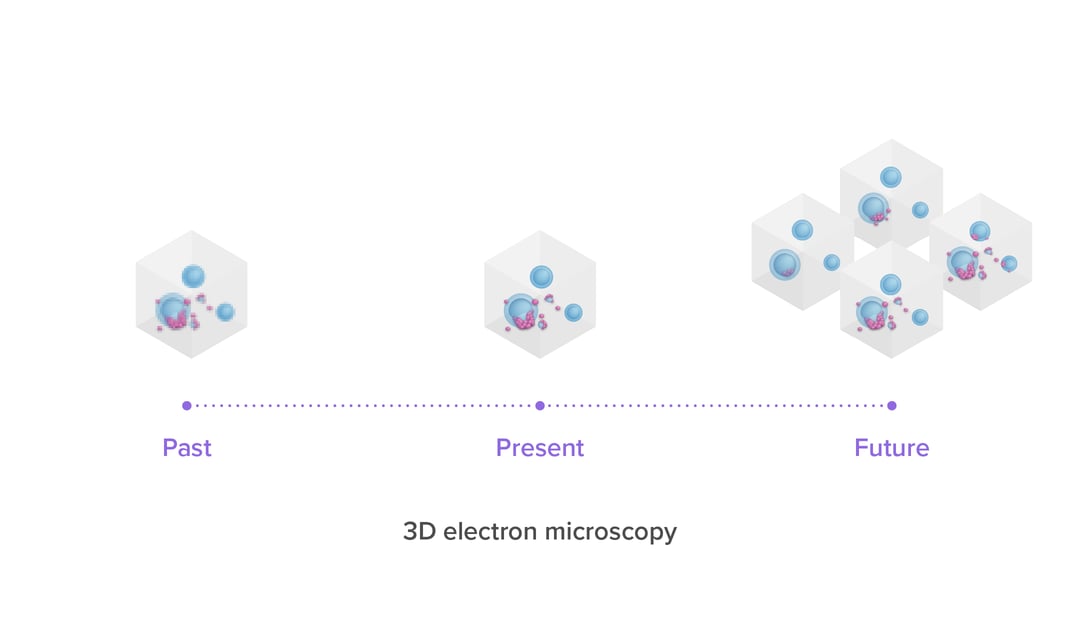
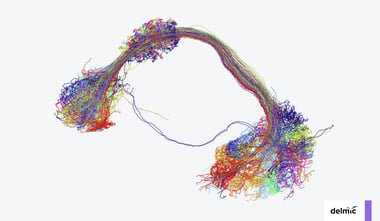
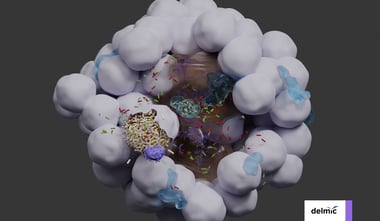
Over the last few decades, the volume capacity and resolving power of 3D electron microscopy (3DEM) has increased substantially, thanks to advances in resolution, automation for volume imaging and data processing power [2,3]. The past 15 years, these advances have enabled 3DEM projects to evolve from spanning small sub-regions of tissue [1] to spanning entire organs at high resolution. As mentioned in our previous blog post, the technique of 3DEM already makes for a perfect tool for imaging and analysis of connectomes. In this blog post we will focus a bit more on the use of 3DEM in cancer research and how the technique helps researchers to understand disease progression, diagnose patients and develop new treatments.
Hallmarks of cancer
Cancer is a heterogenous disease which can acquire multiple hallmarks throughout its multistep development in human tumors. The six main hallmarks are evading growth suppressors, resisting cell death, sustaining proliferative signaling, enabling replicative mortality, activating invasion and metastasis, and inducing angiogenesis (sprouting of new blood vessels) [4]. When looking at both cellular organization and interactions between (cancer) cells and the tumor microenvironment, all these hallmarks can be observed by using high-resolution imaging.
Ultrastructural details in different types of tissues
With the high-resolution capabilities of 3DEM, researchers are looking into all kinds of different applications for this technique. One of such applications is using 3DEM as a diagnostic technique in cancer research. The high resolution of EM is highly useful for detecting minimal changes in cells and tissues, which might not have been seen if light microscopy was used. Current research shows that 3D scanning electron microscopy (SEM) images ultrastructure in a fast and accurate manner [5]. With the use of 3DSEM, different tumorigenesis hallmarks can be identified, enabling distinction between healthy tissue and tumor cells. It is possible to get comprehensive ultrastructural details of several types of tissues, such as renal, brain, prostate and lung tissue, where essential information can be gained to get an accurate pathological diagnosis. This direction for 3DEM application is still under investigation, but in combination with already established techniques it could become a useful tool for both fundamental cancer research and patient diagnosis. In addition, the high throughput of future 3DEM workflows opens the way for comparative studies, where 3DEM volumes from different tumors or biopsies are compared to discover features common during tumorigenesis.
In vivo imaging with 3DEM
One of the most promising next steps for imaging in cancer research, is to not only get high-resolution 3DEM data, but also get more information on how a tumor develops its hallmarks, such as activating invasion and metastasis, over time. In research conducted by Karreman et al. (2015), 3DEM was combined with intravital microscopy (IVM) and microscopic X-ray computed tomography (microCT) to get this high-resolution 3D information of tumor cells and their metastatic properties [6]. The researchers captured single tumor cells from the vasculature of the brain and the skin of mice. The 3DEM images of these cells are correlated with the use of microCT to fit the IVM data and this combination provides the possibility to study the ultrastructure of these cells at a higher resolution and in vivo.
The future of 3DEM
As these research areas show, the application of 3DEM in cancer research is still under development. However, they also indicate that the technique holds great potential, as it has the resolution needed to image biological processes on a nanoscale level. Delmic, together with Thermo Fisher, Technolution and TU Delft, has developed a highly automated 3DEM system, known as FAST-EM. This multibeam electron microscope uses 64 electron beams, allowing for faster, parallel imaging of the sample, decreasing the time needed to image a large 3D sample. With a decreased dwell time per pixel (400ns), the sample faces less beam damage while retaining a high signal-to-noise ratio. Just like other 3DEM techniques, integration of our user-friendly FAST-EM system into a standard workflow will enable high resolution, large volume imaging for cancer research.
Would you like to know how FAST-EM can help to improve your work? Watch our recent launch demonstration to see what FAST-EM can do for you.
References
[1] Denk W, Horstmann H (2004) Serial Block-Face Scanning Electron Microscopy to Reconstruct Three-Dimensional Tissue Nanostructure. PLoS Biol 2(11): e329. https://doi.org/10.1371/journal.pbio.0020329
[2] Hildebrand, D., Cicconet, M., Torres, R. et al. Whole-brain serial-section electron microscopy in larval zebrafish. Nature 545,345–349 (2017). https://doi.org/10.1038/nature22356
[3] Motta, A., Berning, M., Boergens, K. M., Staffler, B., Beining, M., Loomba, S., Hennig, P., Wissler, H., & Helmstaedter, M. (2019). Dense connectomic reconstruction in layer 4 of the somatosensory cortex. Science (New York, N.Y.), 366(6469). https://doi.org/10.1126/science.aay3134
[4] Hanahan, D., & Weinberg, R. A. (2011). Hallmarks of Cancer: The Next Generation. Cell, 144(5), 646–674. https://doi.org/10.1016/j.cell.2011.02.013
[5] Cohen Hyams, T., Mam, K., & Killingsworth, M. C. (2020). Scanning electron microscopy as a new tool for diagnostic pathology and cell biology. Micron, 130, 102797. https://doi.org/10.1016/j.micron.2019.102797
[6] Karreman, M. A., Mercier, L., Schieber, N. L., Solecki, G., Allio, G., Winkler, F., Ruthensteiner, B., Goetz, J. G., & Schwab, Y. (2015). Fast and precise targeting of single tumor cells in vivo by multimodal correlative microscopy. Journal of Cell Science, 129(2), 444–456. https://doi.org/10.1242/jcs.181842
.png)