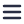
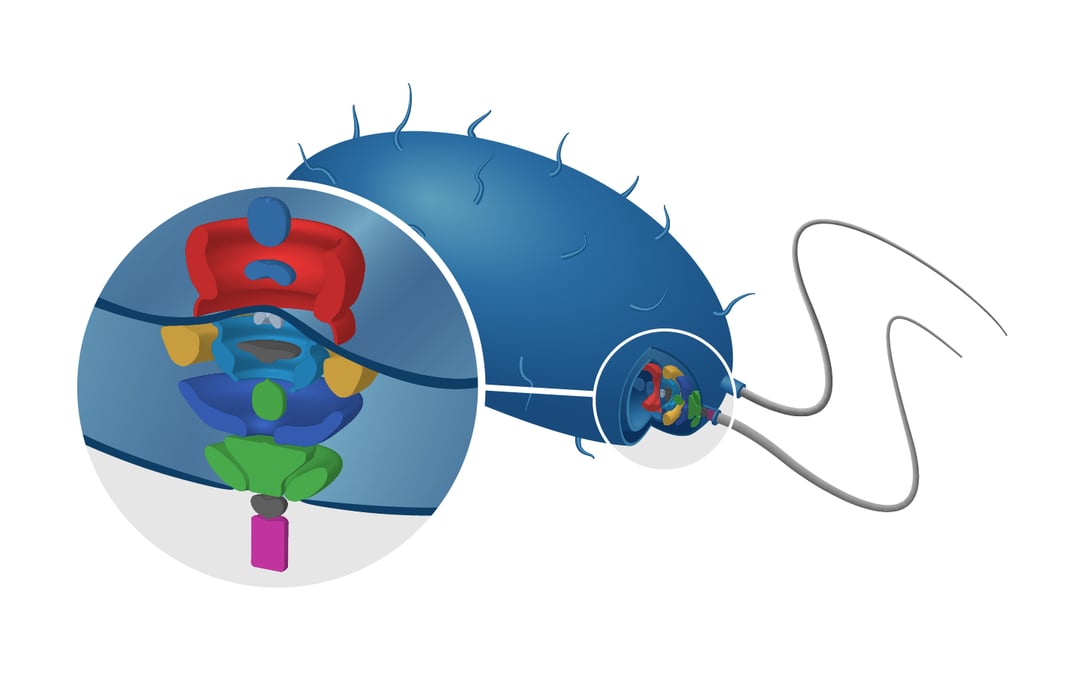
We will discuss the structure of flagella located on the membrane of gamma proteobacteria. These protein systems are what give these organisms the possibility to sense and move to favorable environments.
Determination of flagellar protein structure before 2013
Information about the structure of flagellar proteins is needed to get a better understanding of how these proteins act in their natural environment. In 2012, a review of Stock et al. showed how the combination of various techniques could paint a good picture on the structure of these proteins [1]. At the time, X-ray crystallography and single particle analysis EM were mainly used to resolve the architecture of flagellar motor structures. Although these techniques are powerful tools for protein structure determination, over the last years in situ imaging techniques have taken over.
One of the most powerful imaging tools currently used to resolve proteins in their near-native state is cryo-ET. This technique can image the cellular landscape at near atomic resolution. By tilting the sample, a 3D image can be generated with a resolution as high as 5Å [2]. Recent use of cryo-ET in research on flagellar motor proteins have accelerated the understanding of these complex protein structures.
Imaging the protein architecture
In 2019, Kaplan et al. showed that it is now possible to image the structures of flagellar motors at macromolecular resolution in situ [3]. Using cryo-ET, the researchers imaged the structure of dual-stator systems in γ-proteobacteria with a resolution of up to 10 nm. The 3D reconstructions of the tilt images were calculated through sub-tomogram averaging. The three different stator system architectures were compared to each other and together with bioinformatic homology analysis, the researchers found a correlation between architecture and ion dependency and hypothesize the presence of an evolutionary pathway.
Analyzing chemosensory array evolution
In another study conducted by Ortega et. al., the combination of cryo-ET and bioinformatics also proved to be successful to research the evolutionary origins of the chemosensory machinery that controls flagellar motility in E. coli bacteria [4]. Images were taken of the protein complexes in normal and stress conditions with cryo-ET, after which 3D sub-tomogram averages were computed to get an overview of the different chemosensory architectures in various γ-proteobacterial species. When comparing the architectures in normal and stress condition a change was noticed in the chemosensory array of one of the classes. This effect, imaged by cryo-ET, and data from phylogenetic analysis and homology analysis suggests that consecutive steps, such as loss of components and protein domain swapping, took place in the evolution of the chemosensory array of E. coli bacteria.
The future of Cryo-ET
Cryo-ET has opened up the possibility to study the complex flagellar motor complexes in situ at high resolution. As this powerful technique matures further, we can expect even higher resolution structural data to be obtained. However, Cryo-ET is also a particularly complex technique. At Delmic we design solutions to significantly improve and simplify the cryo-ET workflow.
Check out our previous blog post, introducing METEOR, our integrated cryo-FLM microscope, to get more information on the system and how it can boost your workflow!
References
[1] Stock, D., Namba, K., & Lee, L. K. (2012). Nanorotors and self-assembling macromolecular machines: The torque ring of the bacterial flagellar motor. Current Opinion in Biotechnology, 23(4), 545–554. https://doi.org/10.1016/j.copbio.2012.01.008
[2] Danev, R., Yanagisawa, H., & Kikkawa, M. (2019). Cryo-Electron Microscopy Methodology: Current Aspects and Future Directions. Trends in Biochemical Sciences, 44(10), 837–848. https://doi.org/10.1016/j.tibs.2019.04.008
[3] Kaplan, M., Ghosal, D., Subramanian, P., Oikonomou, C. M., Kjaer, A., Pirbadian, S., Ortega, D. R., Briegel, A., El-Naggar, M. Y., & Jensen, G. J. (2019). The presence and absence of periplasmic rings in bacterial flagellar motors correlates with stator type. ELife, 8. https://doi.org/10.7554/elife.43487
[4] Ortega, D. R., Yang, W., Subramanian, P., Mann, P., Kjær, A., Chen, S., Watts, K. J., Pirbadian, S., Collins, D. A., Kooger, R., Kalyuzhnaya, M. G., Ringgaard, S., Briegel, A., & Jensen, G. J. (2020). Repurposing a chemosensory macromolecular machine. Nature Communications, 11(1). https://doi.org/10.1038/s41467-020-15736-5
.png)