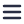
Organelles require constant interaction with other parts of the cell. This is achieved via a network of membrane enclosed vesicles and tubules that shuttle between all organelles and thereby connect all parts of the cell. The dynamic process of vesicular trafficking and membrane contacts between organelles is called membrane trafficking.
Membrane trafficking is fast and very dynamic. Because of this, live-cell fluorescence microscopy is the preferred method to study the trafficking pathways of the cell. However, many structures of the membrane trafficking process, such as endocytic vesicles and membrane folds, are smaller than the resolving power of light microscopes. Therefore, Electron Microscopy is used to get high resolution information about the compartments in the cell. Studying the morphology of organelles, vesicles and tubules using Transmission Electron Microscopy (TEM) laid the foundation of our current understanding of the organization of the cell.
TEM is performed using thin sections of cells. Because of this, organelles are always viewed as a projection of a thin slice. However, organelles and vesicles have very complex 3D shapes that are closely tied to their function. To extract 3D information is very difficult with TEM. This is why Electron Tomography has become a valuable tool to gain 3D structural information on membrane compartments. Electron Tomography has been instrumental for the study of several biological aspects of the cell (such as autophagosome formation [1] and the 3D structure of the Golgi [2] and Endoplasmic Reticulum [3]) that were overlooked with TEM.
Taking it one step further, with cryogenic electron tomography (cryo-ET) it is possible to acquire high-resolution 3D information at cryogenic temperatures. Imaging at low temperatures preserves the morphology of the membrane compartments and protein complexes in the cell. Therefore, structural information on protein complexes that reside on membranes can be acquired to give additional information on the compartments in question. This combination of a high resolution with near-native ultrastructural morphology has opened the door for new insights in the field of membrane trafficking. Where traditionally protein complexes such as Clathrin and Ribosomes could only be seen in enough detail to identify specific membranes, with cryo-ET these protein complexes can be resolved and studied in detail on their native membrane [4].
Cryo-ET holds the promise to take the field of membrane trafficking a step further by combining 3D high-resolution near-native morphological detail of intracellular membranes with structural information of membrane-resident protein complexes.
References
[1] M. Hayashi-Nishino, N. Fujita, T. Noda, A. Yamaguchi, T. Yoshimori, and A. Yamamoto, “A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation,” Nat. Cell Biol., vol. 11, no. 12, pp. 1433–1437, Dec. 2009, doi: 10.1038/ncb1991.
[2] M. S. Ladinsky, J. R. Kremer, P. S. Furcinitti, J. R. McIntosh, and K. E. Howell, “HVEM tomography of the trans-Golgi network: Structural insights and identification of a lace-like vesicle coat,” J. Cell Biol., vol. 127, no. 1, pp. 29–38, Oct. 1994, doi: 10.1083/jcb.127.1.29.
[3] M. E. Martone, Y. Zhang, V. M. Simpliciano, B. O. Carragher, and M. H. Ellisman, “Three-dimensional visualization of the smooth endoplasmic reticulum in Purkinje cell dendrites,” J. Neurosci., vol. 13, no. 11, pp. 4636–4646, 1993, doi: 10.1523/jneurosci.13-11-04636.1993.
[4] F. Brandt, L. A. Carlson, F. U. Hartl, W. Baumeister, and K. Grünewald, “The Three-Dimensional Organization of Polyribosomes in Intact Human Cells,” Mol. Cell, vol. 39, no. 4, pp. 560–569, Aug. 2010, doi: 10.1016/j.molcel.2010.08.003.
This work is supported by the European SME2 grant № 879673 - Cryo-SECOM Workflow
.png)